Using the mfold Web Server With Constraints
- 1. Forcing a string of consecutive bases to pair.
- This is an interesting constraint. The partners for these bases are chosen by the program. To enter this type of constraint, type:
F i 0 k
on a line in the constraint box. The letter "F" means "Force"; the first number, "i", is the first base that must pair; the second number, "0" (zero), is not used, and the third number, "k", is the number of consecutive bases that must pair, beginning with the ith. For example,
F 23 0 5
would cause bases 23, 24, 25, 26 and 27 to pair. - 2. Forcing a string of consecutive base pairs.
- This is the same thing as forcing a helix to form. The helix is designated by its external closing base pair. That is, by the base pair i.j where iis as small as possible and j is as large as possible. Thus the k consecutive base pairs beginning with i.j are: i.j, i+1.j-1, i+2.j-2, ..., i+k-1.j-k+1. To enter this type of constraint, type
F i j k
on a line in the constraint box. Note how 3 integers are necessary in this case. For example,
F 2 110 3
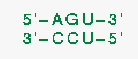 would force the following base pairs: 2.110, 3.109, 4.108. These base pairs must be able to formBe aware also that mfold filters out isolated base pairs. In the example on the right, the G.C base pair is isolated.
would force the following base pairs: 2.110, 3.109, 4.108. These base pairs must be able to formBe aware also that mfold filters out isolated base pairs. In the example on the right, the G.C base pair is isolated. - 3. Prohibiting a string of consecutive bases from pairing.
-
To enter this simple constraint, type:
P i 0 k
on a line in the constraint box. This command is similar to 1. above, except that the designated bases are prevented from pairing.
- 4. Prohibiting a string of consecutive base pairs
-
To enter this simple constraint, type:
P i j k
on a line in the constraint box. This command is similar to 2. above, except that the designated base pairs are prevented from forming.
- 5. Prohibiting 1 part of a sequence from pairing with another
- If your molecule has N bases, and 1 <= i <= j <= N & 1 <= k <= l <= N, are 2 segments, then these segments can be prevented from pairing with each other by entering
P i-j k-l
on 1 line in the constraint box. For example, in a molecule with 120 bases, the constraint
P 1-10 11-120
will prevent the first 10 bases from pairing in any way with the last 10 bases. The constraint
P i-j i-j
will prevent the segment i-j from pairing with itself. - 6. Annotated bases
- The software recognizes A, C, G, U and T. In RNA folding, a 'T' will be converted to a 'U'; vice versa for DNA folding. In addition B, D, H and V are recognized as A, C, G and U/T, respectively. Bases marked in this way are regarded as susceptible to nuclease cleavage. They are allowed to pair only if their 3' neighbor is unpaired. This is an old feature of mfold.
The mfold web server also recognizes W, X, Y and Z as A, C, G and U/T, respectively. These bases are regarded as "modified" and are allowed to pair only at the ends of helices. Please note that the commonly used ambiguous codes shown in the table below ARE NOT SUPPORTED by the mfold server.
Unsupported ambiguous codes for DNA: A,G
RC,T
YA,T
WC,G
SA,C
MG,T
KC,G,T
BA,G,T
DA,C,T
HA,C,G
VA,C,G,T
N
