Example 1
Next: EXAMPLE 2 Up: Sample foldings Previous: Sample foldings
EXAMPLE 1
The energy dot plot is an integral part of the folding prediction. Consider
the folding of a short RNA sequence:
ACCCCCUCCU UCCUUGGAUC AAGGGGCUCA A,
using default parameters. ΔG = -9.8 kcal/mol at 37°, so ΔΔG = 1.0 rather than 5% of ΔG. A single, optimal folding is
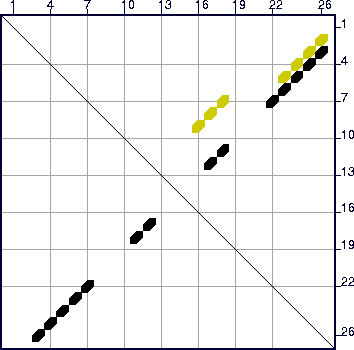
computed. A glance of the energy dot plot , shown in Figure 13, reveals the optimal folding in black dots (symbols), but
another set of yellow dots, indicating base pairs in at least 1 other suboptimal
folding. The default value of `W' (2, from Table 3) is too large
for this other folding to be predicted, but a glance at the dot plot shows that
something else is there. When the sequence is refolded with `W'=0, a second, totally
different folding is predicted. Figure 14 displays these
foldings with individual bases drawn.
 |
![\begin{figure}\centering \subfigure[]{\epsfig{file=1-again_1_zoom.ps,width=0.3\t... ...\subfigure[]{\epsfig{file=1-again_2_zoom.ps,width=0.3\textwidth} }\end{figure}](img123.gif) |
Next: EXAMPLE 2 Up: Sample foldings Previous: Sample foldings
 |
Michael Zuker Center for Computational Biology Washington University in St. Louis 1998-12-05 |